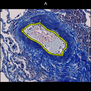
Evaluation Data for Virtual Multistaining Registration
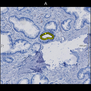
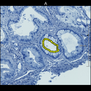
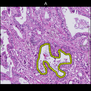
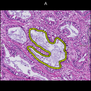
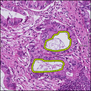
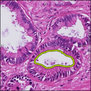
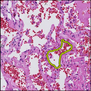
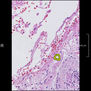
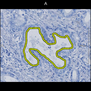
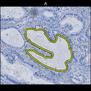
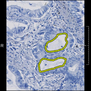
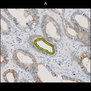
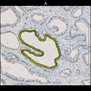
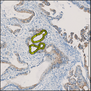
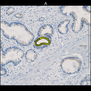
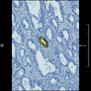
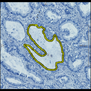
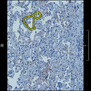
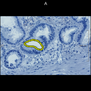
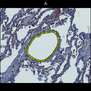
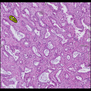
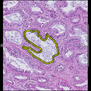
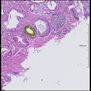
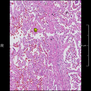
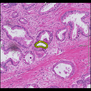
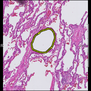
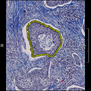
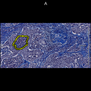
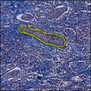
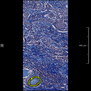
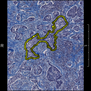
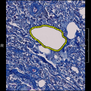
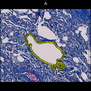
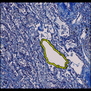
Comparative evaluation of registration methods applied to histology data is difficult due to the lack of ground truth data in most datasets. We provide annotated histology data of multiple stains that can be used for registration. The accuracy of the registration can be evaluated based on corresponding structures that have been segmented manually.
Data description
The image data is available as Hamamatsu (ndpi) files and annotations are available as point lists in a sqlite-database format. A sample python file is provided to access the annotations.
Annotations
We manually segmented between five and twelve structures in each image. The points of the segmentations are stored in files annotations/[image_file_name].sqlite. The simplest way to access this data is by using the sample python file provied. You may limit the x- and y- ranges to only select a specific structures. Currently, the individual structures can not be separated solely based on the sqlite data (we are working on it).
Download: annotations.zip readAnnotations.py
Image data
Currently, ten slide pairs (consisting of 13 slide images) are available from three different tissue blocks:
Block L0:
L0_001-847 10_D2 2_01_CD31 - 2012-06-19 11.17.42.ndpi
L0_002-847_10_d2_2_he_2_-_2012-05-22_11.52.54.ndpi
L0_003-847-10_D2,2_3_-_2012-08-31_14.21.38.ndpi
L0_004-4_-_2012-09-20_12.04.09.ndpi
L0_005-847_10_D2 2_05_CD31_-_2012-06-19_11.27.00.ndpi
L0_006-847_10_d2_2_he_6_-_2012-05-22_11.57.28.ndpi
Block L1
L1_002 - 2014-05-27 12.12.29.ndpi
L1_003 - 2014-08-19 09.16.01.ndpi
L1_004 - 2014-05-27 12.16.14.ndpi
L1_005 - 2014-08-19 09.19.27.ndpi
L1_006 - 2014-05-27 12.19.57.ndpi
Block K1
LL1_1_CD146 - 2014-12-12 12.09.18.ndpi
LL1_2_AFOG - 2014-12-12 13.00.40.ndpi
The image data is available for download upon request only. Please contact Johannes Lotz or Nick Weiss to get access.
Acknowledgment
We thank the following people for their contributions to these datasets:
Institute of Pathology, University Hospital Heidelberg
- Kai Breuhahn
- Benedikt Müller
- Margarita. González-Vallinas
- Arne Warth
Tissue Imaging and Analysis Center, University of Heidelberg
- Bernd Lahrmann
- Niels Grabe
Fraunhofer MEVIS, Lübeck / Bremen and Institute of Mathematics and Image Computing
- Johannes Lotz
- Janine Olesch
- Nick Weiss
- Judith Lotz
- Patricia Galuschka
- Thomas Polzin
- Hendrik Laue
- Jan Modersitzki
Department of Radiology, German Cancer Research Center and with Diagnostic and Interventional Radiology, University Medical Center Heidelberg
- Oliver Sedlaczek
Part of this work was supported by the MedSys-Network LungSys which is funded by the German Federal Ministry of Education and Research, grant number 0316042J, 0316042B. Tissue samples were provided by the tissue bank of the National Center of Tumor Diseases (NCT, Heidelberg, Germany) and the biobank platform of the German Center for Lung Research (DZL) in accordance with the regulations of the tissue bank and the approval of the Ethics Committee of the Heidelberg University.

- People
- Johannes Bostelmann
- Daniel Budelmann
- Bernd Fischer
- Florian Galow
- Ole Gildemeister
- Stephanie Häger
- Stefan Heldmann
- Temke Kohlbrandt
- Sven Kuckertz
- Annkristin Lange
- Jan Lellmann
- Tanja Loßau
- Johannes Lotz
- Publications
- 3D Histology Reconstructions
- Evaluation Data for Virtual Multistaining Registration
- Jan Modersitzki
- Till Nicke
- Josephine Oettinger
- Nils Papenberg
- Sofie Saier
- Kerstin Sietas
- Angela Tesdorpf
- Herbert Thiele
- Johannes Voigts
- Sina Walluscheck
- Nick Weiss
- Sonja Wichelmann